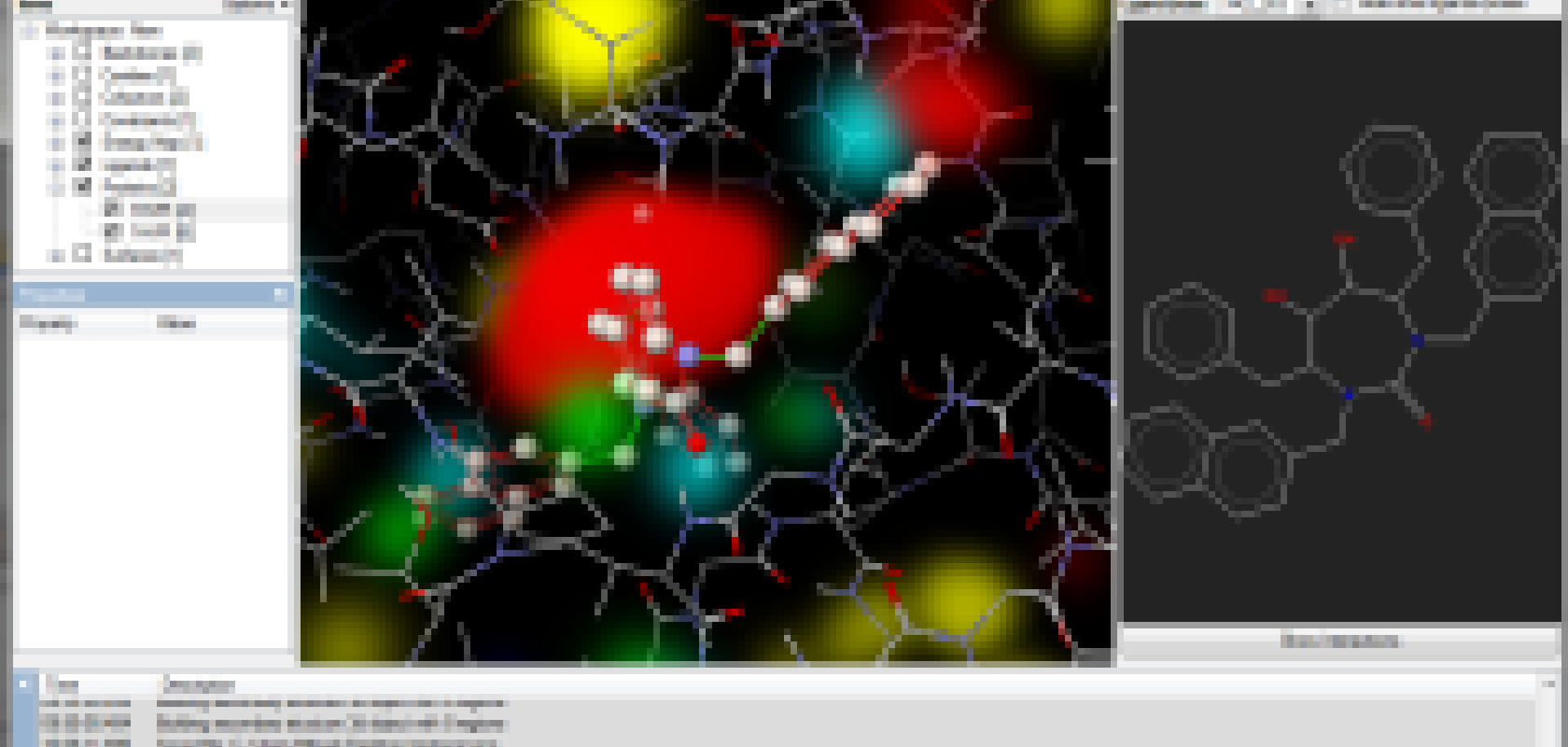
Version 5.5 of Molegro Virtual Docker, an integrated platform for computational drug design available for Windows, Linux and Mac OS X, has been released by CLC bio. Molegro Virtual Docker offers high-quality protein-ligand docking based on novel optimisation techniques combined with a user interface experience focusing on usability and productivity.
New features include a new 'Energy Maps' tool that provides volumetric visualisation of protein force fields; an improved ray-tracer that more closely matches the live 3D view output, enabling the creation of high-resolution renderings of the 3D view; and a new execution mode in the Docking Wizard: 'Run Docking in Multiple Processes'. This enables medium-sized jobs to be run on a local machine, while utilising multiple CPU cores and even multiple GPU graphics cards. The company advises that for large jobs on multiple machines, Molegro Virtual Grid should still be used.