Advances in computer vision combined with AI computing are helping pathologists to more accurately identify subtypes of cancer - leading to better treatments for patients, explains Dan Ruderman.
‘80 years on pathologists still rely on their eyes to diagnose,’ explains Dan Ruderman, Assistant Professor of Research Medicine at the Keck School of Medicine of USC and the Lawrence J Ellison Institute for Transformative Medicine.
‘Pathologists use their eyes to make these decisions so whether they have gone from a microscope to now looking at digital images that have been scanned in from slides, they are still looking at them with their eyes and making those decisions’ Ruderman continues.
Ruderman and his colleagues at the Ellison Institute have been using large scale AI simulations run in partnership with Oracle using the Oracle Cloud Infrastructure. Their research looks to take pre-existing Hematoxylin and eosin stain (H&E stains) and using them to identify cancer subtypes by training neural networks.
‘Every patient around the world who goes through surgery and has a pathologist examine specimens has an H&E stain done to find out what the diagnosis of this patient is,’ states Ruderman. ‘This means that it costs zero dollars because the test has already been done.’
Using computers to assist in diagnosis
‘What is surprising about that is given the amazing advances we have had in computer vision for things like self-driving cars and certain modalities in medical imaging that we are not using the advanced capabilities of computers to help in these diagnoses to make better decisions for patient care.’
Today the fields of visual pathology and AI and machine learning are being employed to solve particularly challenging questions in the diagnosis of cancer subtypes. This is an important step in the effective treatment of cancer because it means that patients can potentially receive the correct treatment faster and without having unwanted side effects from drugs that may be ineffective.
‘The big question in clinical care is who is going to respond to which therapy. Once a diagnosis has been made what subtype of, in my case, cancer is it which determines the kind of therapies that a patient is going to get.’
There are a number of existing molecular markers that can help to identify potential treatments but while ‘they are good, they are not good enough,’ argues Ruderman. ‘For example, if there is a mutation in a patient's melanoma to the BRAF gene then they would be given Zelburaf as the drug to treat them. But despite having this molecular marker only around half of those patients are going to respond to that drug,’ added Ruderman. ‘What is common to all of these is that patients are going to have to suffer the side effects of these therapies but not achieve any benefit.
The question that researchers at the Ellison Institute began to ask themselves is: ‘can AI help to make better decisions in these cases to predict, not just which patients have these markers but which patients will respond?’
The initial research looked at hormone receptor-positive breast cancer. Traditionally this is done with a $300 immuno-histochemistry test, if a sample is found to be estrogen receptor-positive (ER+) then approximately 60 per cent of patients respond to endocrine therapy such as Tamoxifen.
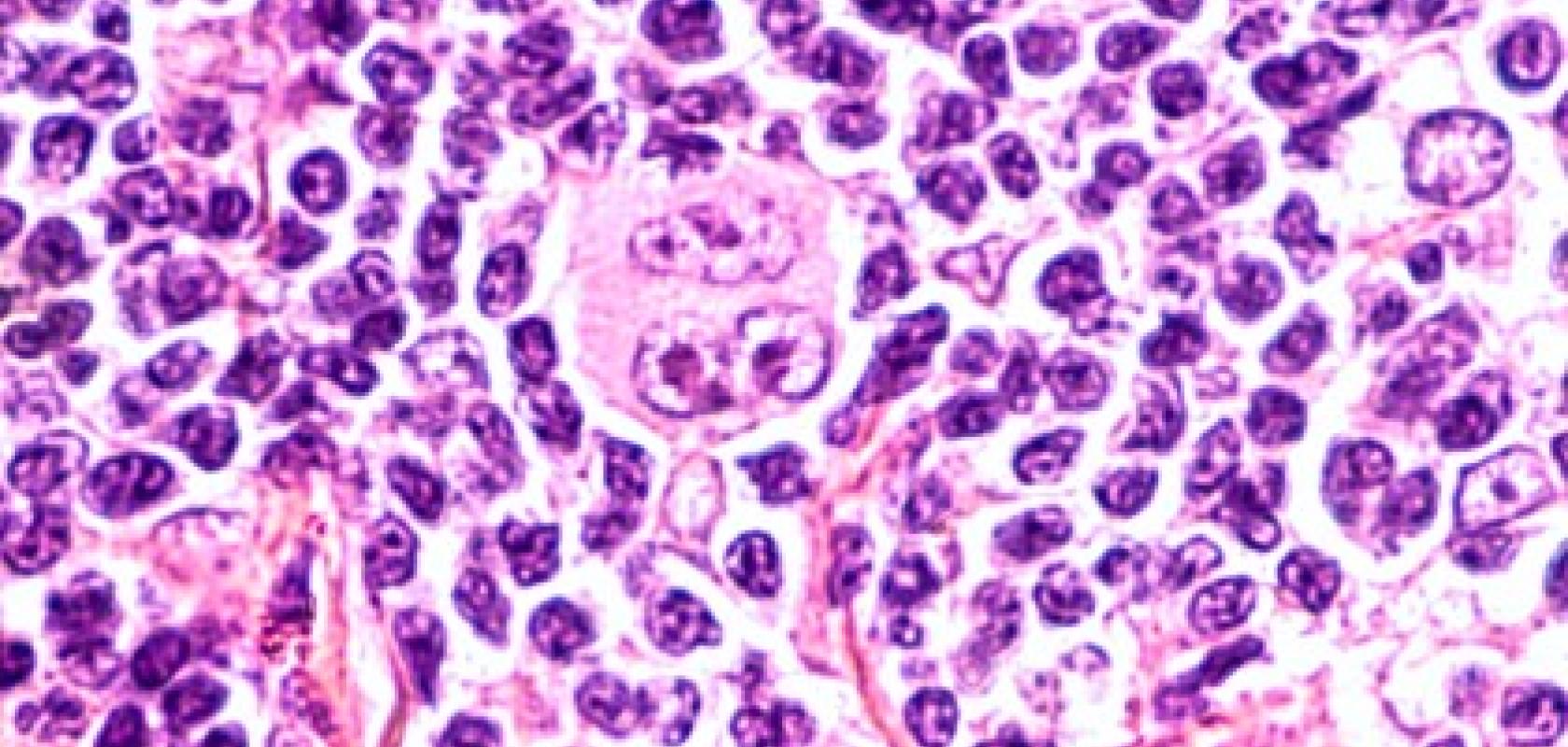
‘Fundamentally these two cancers look the same under the microscope. If you ask a pathologist to designate just by eye from this kind of scan - is it an ER+ or ER- tumour? They generally would not be able to do it,’ stated Ruderman. ‘There are certain subtypes such as lobular cancers where the cells all arrange in lines which could be identified as ER+ cancer but that is a fairly rare subtype.
‘Can we use AI and all the wonderful advances in computer vision to “read the tea leaves” and find things that are too subtle for our eyes to discern?’
The slides are digitised and then broken down into different measurable variables such as the size of nuclei, orientation etc this is fed into a deep (more than five layers) neural network.
Testing on non trained samples enabled the researchers to create an ROC curve, which is a graphical plot that illustrates the diagnostic ability of a binary classifier system. The initial area under the curve (AOC) was found to be 0.72.
While initial results were promising several iterations of the network and advances in organisation, compression and fingerprint technologies have meant that the accuracy of prediction using the H&E stain has risen dramatically. AUC in the later tests was found to be 0.89 which provides a much better standard for diagnosis.
Running on the Oracle Cloud Infrastructure, researchers at the Ellison Institute bare-metal GPU instances that are running and training these Deep convolutional neural networks. They are also making use of Oracle Autonomous Database to store a lot of their training and inference data. ‘We have tens of millions of rows in these database tables,’ comments Ruderman. ‘Then on the shared file system, we have the whole slide images. Terabytes of data from all of these slides that have data captured.’
The hope for the future is to take multiple slides and produce a 3D model which could provide much greater accuracy as the amount of data contained in each patient sample would increase dramatically.
Ruderman also hopes that the work done here and at other research centres looking at different cancer types or other diagnosis can help to develop the frameworks for wider research into this combination of medicine, AI and computing vision.
Looking to the future
‘We can take that exact same slide that has already been created and extract more information out of it. This information can tell a clinician that they may benefit from hormonal therapy and that is valuable because those pills are cheap, for example,’ said Ruderman. ‘It is information that is already held in the slides they have in their hands. You just need to get it digitised and into a computer so they can make the analysis.’
This data could be collected at the point, or it could be mailed off and scanned at another location. However Ruderman stressed that this should not be seen as a replacement for traditional methods, rather it should be used to supplement and improve doctors ability to diagnose.
‘For every cancer, there is an alphabet soup of these kinds of mutations or tests that you can do to subclassify these according to what therapies they will respond to. If you look at the publications in digital pathology people are working on lung cancer, colorectal cancer, prostate cancer and others,’ comments Ruderman.
‘They are not exactly asking the same questions that we are asking but people are now digitising those samples and running deep convolutional neural networks on them.’
In this new age of AI and deep learning, Ruderman stressed that there is no magic bullet to develop the perfect model. ‘We are still in the age where it is know-how, trial and error and an art form. The more computing power you have the more room you have to experiment and try different architectures simultaneously’ said Ruderman.
This is a key part of the collaboration between the Ellison Institute and Oracle as the computing power provided By Oracle allows the researchers to try different model parameters to explore the ways that the model can be improved.
‘People are now getting things to work at a certain level and they publish their architecture. You might look at the number of layers and try to emulate it or build from that existing process or protocol that has been developed previously.
‘That is where having a lot of computing resources that we have with our collaboration with Oracle, is really helpful because it allows us to explore these things. The short answer is, there is no formula, go out for yourself and try stuff,’ stated Ruderman.
‘You take something that works, start there and then you tweak it. That is the world we are in right now. As computing power becomes more accessible that means we can experiment more,’ Ruderman concluded.